Plant Disease Recognition System
End-to-end ML pipeline for leaf disease classification: a CNN that classifies plant leaf images into 38 disease/healthy classes, with a single config, shared data pipeline, and a Streamlit web app—built for reproducibility and train/serve consistency.

In a nutshell
- 38-way classification across 14 crops (e.g. Apple, Corn, Tomato, Potato, Grape)—healthy and disease states—trained on ~87k images (70% train / 15% val / 15% test).
- Single source of truth:
config.pyholds paths, hyperparameters, seed, and the 38 class names in fixed order;data_pipeline.pydefines preprocessing and augmentation so training and the app never drift. - Thin Streamlit app: user uploads a leaf image → resize to 128×128, normalize the
same way as training → model predicts → label via
config.CLASS_NAMES. Model is cached; app detects Rescaling layer and feeds [0,1] or [0,255] so inference matches the trained model. - Reproducibility: seed control, experiment logs (history + metrics + config
snapshot) under
experiments/; CI with Ruff lint and smoke tests on GitHub Actions; deployment documented for Streamlit Community Cloud.
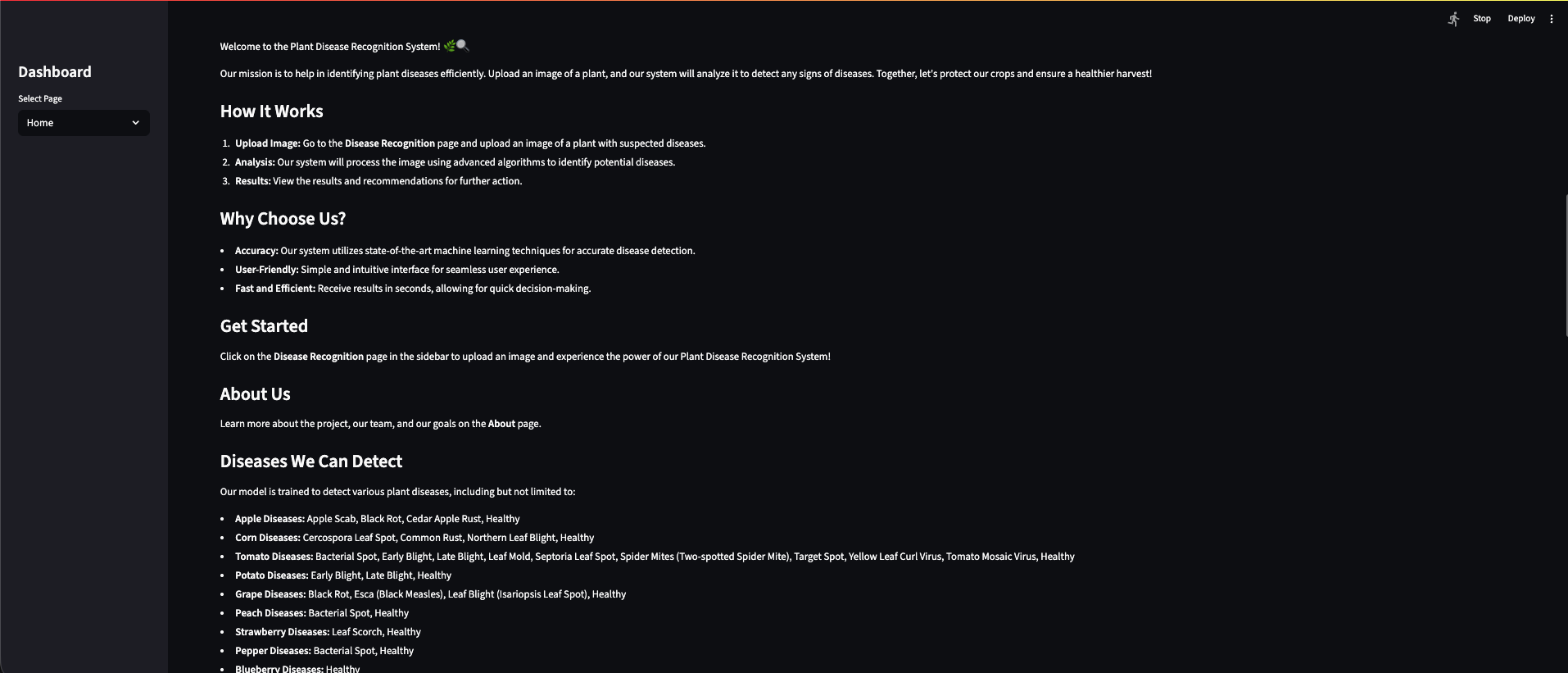
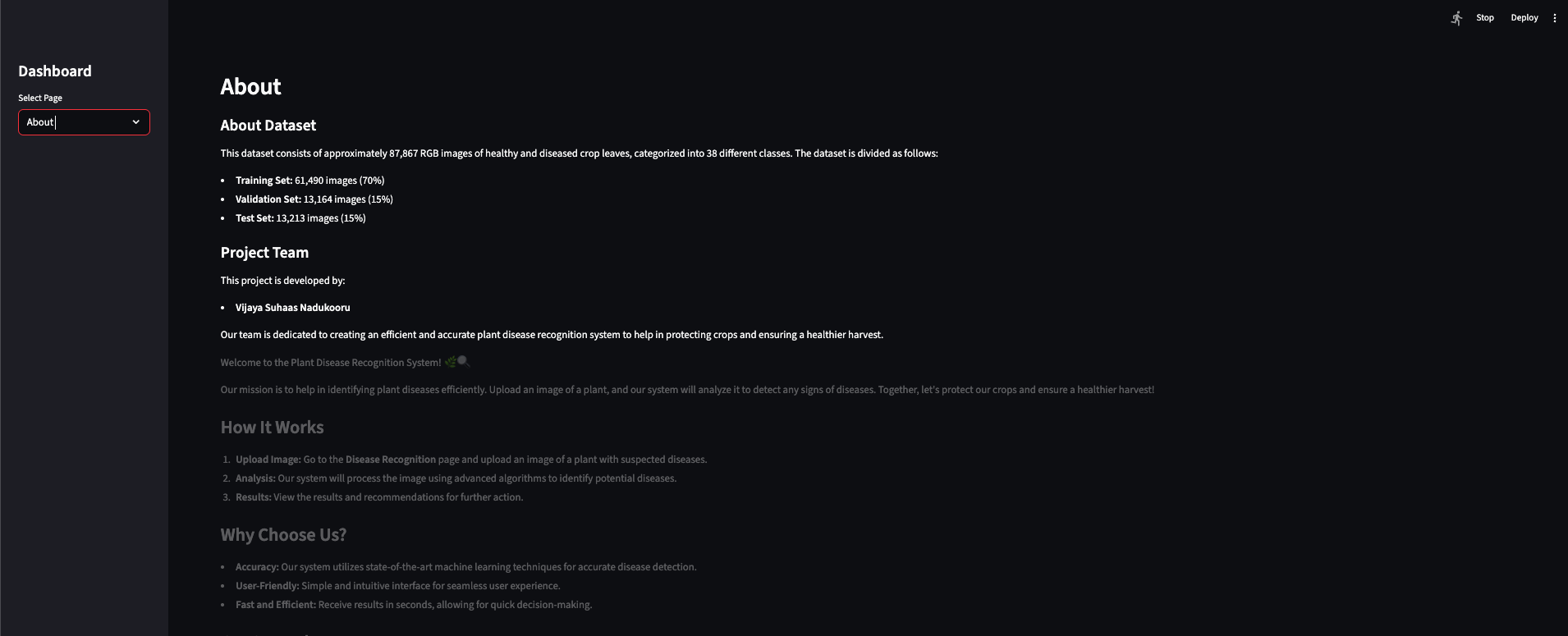
In the app
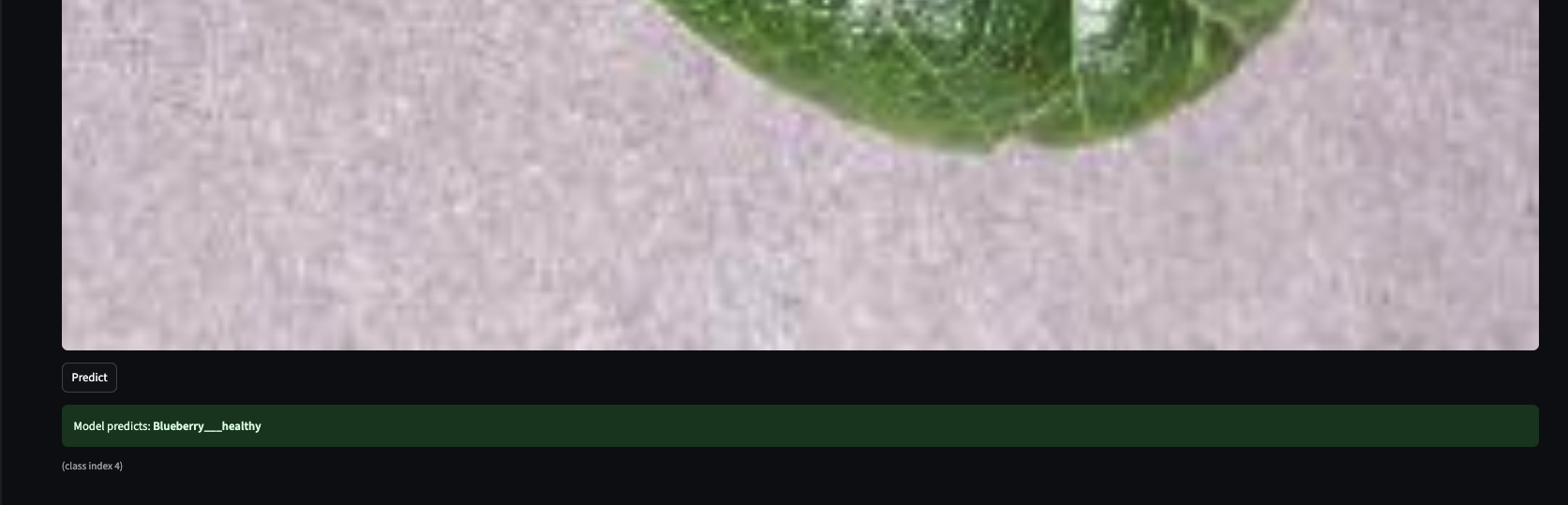
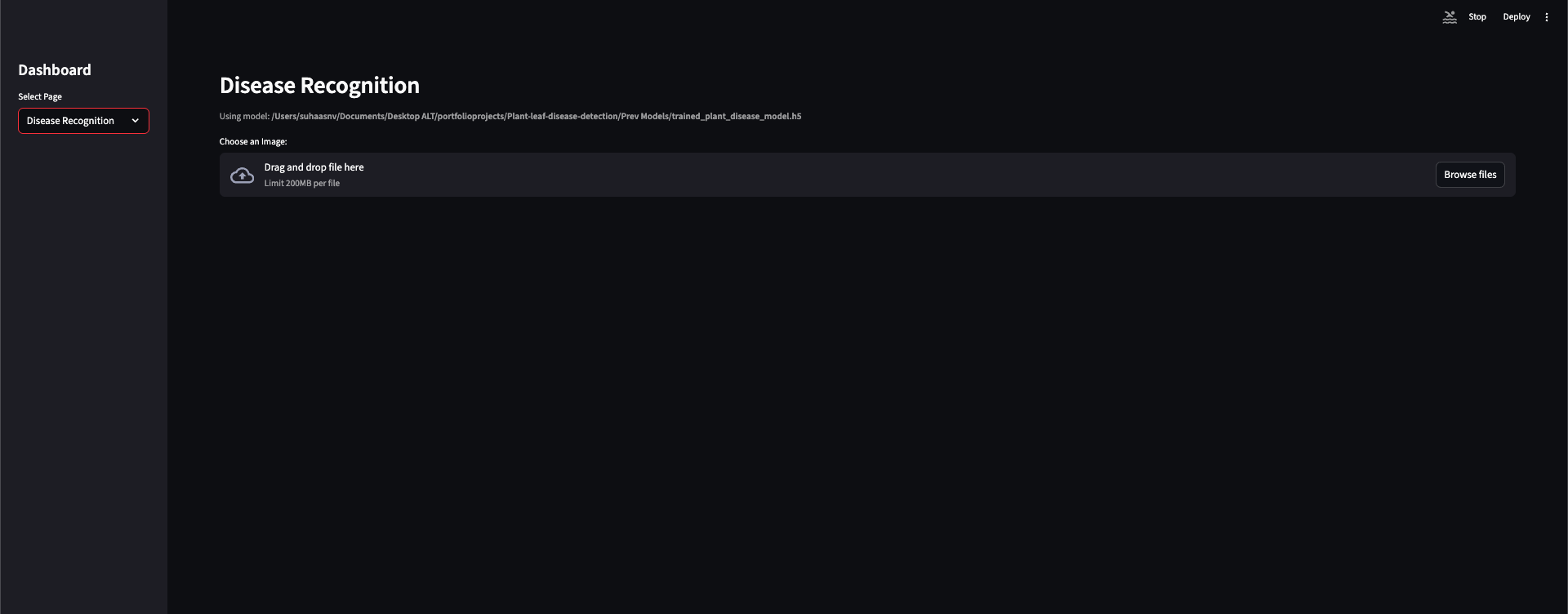
Home and About introduce the system; Disease Recognition is where you upload a leaf image and get a classification.

What it is
A deep learning project that detects plant diseases from leaf images—38 classes, ~95% accuracy—with a centralized config and data pipeline, and a Streamlit app that stays in sync with training.
The Plant Disease Recognition System is an end-to-end ML system: you point it at a leaf image and it tells you whether the plant is healthy or which disease it has. Under the hood, a CNN (TensorFlow/Keras) classifies into 38 labels—covering 14 crops and their healthy/disease variants—trained on a directory-based image dataset split into train, validation, and test.
What I cared about from the start was not duplicating logic. It’s easy to have one
preprocessing path in a Jupyter notebook and another in the app, or to change class order and break
predictions. So I built a single config.py (paths, image size, batch size, learning rate,
epochs, seed, augmentation settings, and the 38 class names in a fixed order) and a single
data_pipeline.py that both the training notebook and the Streamlit app rely on. Training
and serving see the same preprocessing and the same labels—no train/serve mismatch, no “always predicts
one class” surprises.
How it works
Data lives in folders per class; config and pipeline drive everything; one training path produces the model; the app loads it once, preprocesses the same way, and shows the label.
Leaf images sit in data/train, data/valid, and data/test—one
folder per class. Paths and split logic come from config.py. The data
pipeline handles resize (e.g. 128×128), normalization to [0,1], and augmentation (flips,
rotation, zoom, brightness) only for training. It builds train/val/test datasets and an inference
preprocessing path so that whatever the model saw during training is exactly what the app feeds at
prediction time.
In the training notebook I set the seed, build datasets from the pipeline, define a CNN
(Conv blocks, dropout, dense head, 38-way softmax), train with EarlyStopping and ReduceLROnPlateau,
evaluate on train/val/test, and save history and metrics to experiments/ plus the model
(e.g. .h5) in the project root or a Prev Models/ folder. The saved Keras model
is then loaded once in the Streamlit app (cached). The app checks whether the model has a Rescaling
layer and normalizes input to [0,1] or [0,255] accordingly—so inference always matches how the model was
trained.
What it does for users
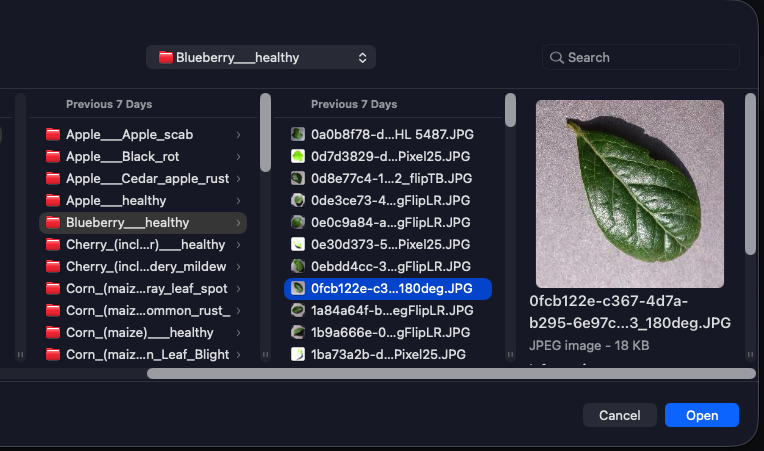
Upload a leaf image in the Streamlit app; get an instant classification (healthy or disease) with the correct label from the same 38 classes the model was trained on.
The Streamlit app is deliberately thin: the user uploads an image, the app resizes it to
the configured size (128×128), normalizes it the same way as in the pipeline, runs
model.predict, and maps the predicted index to config.CLASS_NAMES. The UI can
show which model file is loaded (e.g. from project root or Prev Models/), so it’s clear
what’s in use. No extra features—just upload, predict, and show the label. All ML assumptions (size,
normalization, class order) come from config and the pipeline, so the app never “drifts” from training.
Tech stack
Python 3.9+, TensorFlow 2.x/Keras for the CNN and data pipeline; NumPy, PIL, tf.data; Streamlit for the UI; single config and pipeline; Jupyter for training; GitHub Actions for CI; Streamlit Community Cloud for deployment.
Language & ML: Python 3.9+, TensorFlow 2.x, Keras—CNN, image preprocessing, data
augmentation. Data: NumPy, PIL/Pillow, tf.data pipelines, directory-based image
datasets. App: Streamlit (web UI, file upload, model caching). Config &
pipeline: config.py (paths, hyperparameters, seed, class names),
data_pipeline.py (preprocessing, augmentation, train/val/test and inference helpers).
Notebooks: Jupyter for training, evaluation, and saving model and experiment artifacts.
CI/CD: GitHub Actions (Ruff lint, smoke tests); deployment documented for Streamlit
Community Cloud. Version control: Git, .gitignore for data, models, experiments, venv.
Standout technical choices
One config and one data pipeline for training and inference; app adapts to model (Rescaling or not); experiment logging and seed for reproducibility; CI and deployment docs.
Train/serve alignment. The biggest risk in a small ML project is training with one
preprocessing path and serving with another—or changing class order and getting wrong labels. By having
a single config.py and data_pipeline.py imported by both the notebook and the
app, preprocessing and class order stay consistent. The app also inspects the loaded model: if it has a
Rescaling layer, we feed [0,255]; otherwise we normalize to [0,1] before calling predict. That way
different saved models (with or without built-in rescaling) still get the right input range.
Fixing “always one class” and wrong predictions. In practice I ran into wrong or stuck
predictions. The fixes were: (1) using the correct model file (e.g. from Prev Models/), (2)
fixing file stream handling (e.g. seek(0) on uploads so the image is read correctly), and
(3) matching input range to the model—raw [0,255] vs [0,1] depending on whether the model has Rescaling.
Documenting this in the repo helps anyone reusing the app.
Reproducibility. Seed is set in config and used in the notebook; each run can dump a
config snapshot and metrics into experiments/. That makes it easy to compare runs and to
know exactly what hyperparameters and class order a given checkpoint used.
CI and deployment. GitHub Actions runs Ruff on the core Python files and a simple
import/smoke test so the pipeline and app stay runnable. Deployment is documented for Streamlit
Community Cloud (connect repo, set main.py as entrypoint, handle model via repo or external
URL and secrets).
Results
38 classes, ~87k images, 70/15/15 split; CNN with Conv blocks, dropout, dense head; ~95% accuracy on test; Streamlit app with CI and deployment docs.
Classes: 38 (e.g. Apple, Corn, Tomato, Potato, Grape—healthy and disease states). Dataset: ~87k images, 70% train / 15% val / 15% test. Model: CNN (Conv2D blocks, dropout, dense head, 38-way softmax). Accuracy: ~95% on the test set (you can replace this with your exact number in the repo). Deployment: Streamlit app; CI on GitHub Actions; deployment steps documented for Streamlit Cloud.
What I implemented
Full pipeline: centralized config and data pipeline, TensorFlow/Keras training notebook with experiment logging, Streamlit app with correct preprocessing and model caching, CI (GitHub Actions), and deployment documentation.
I designed and implemented the full pipeline: config (config.py with paths,
image size, batch sizes, learning rate, epochs, seed, augmentation settings, and the 38 class names in
fixed order). Data pipeline (data_pipeline.py with preprocessing,
augmentation, dataset builders for train/val/test, and inference preprocessing).
Training (Jupyter notebook: seed, dataset build, CNN definition, EarlyStopping,
ReduceLROnPlateau, evaluation, saving history/metrics to experiments/, saving the model).
App (Streamlit: load model once with caching, detect Rescaling layer, resize and
normalize uploads, predict, display label via config.CLASS_NAMES). CI
(GitHub Actions: install deps, Ruff lint, smoke test). Docs (deployment for Streamlit
Community Cloud, optional script like split_valid_to_test.py for splitting validation into
test). The goal throughout was a single source of truth for paths, hyperparameters, and class labels so
that training and inference stay aligned and reproducible.
What I learned
Train/serve mismatch bites; one config and one pipeline fix it. Input range and file handling matter as much as model architecture. Reproducibility pays off when iterating.
Keeping training and serving in sync is non-negotiable. As soon as you have two places that preprocess or
two copies of class names, something will drift—wrong labels or a model that seems to “always predict
one class.” One config and one data pipeline, both imported by the notebook and the app, removed that
class of bugs. I also learned to always check input range: models with a Rescaling
layer expect [0,255]; those without expect [0,1]. The app has to match, or predictions are off. File
handling matters too—e.g. seek(0) on uploaded streams so the image bytes are read
correctly.
Reproducibility—seed, experiment logs, and a config snapshot per run—made it easy to compare checkpoints and to remember what each saved model assumed. For a portfolio project, that’s a good habit: it shows you care about the full loop, not just the accuracy number.
System Architecture
Training and inference share a single config and data pipeline — the same preprocessing path that builds train/val/test datasets is the path the Streamlit app uses at prediction time. This eliminates the most common class of train/serve mismatch bugs.
ML pipeline — config.py and data_pipeline.py are
the shared core. Both the training notebook and the Streamlit app import them, ensuring preprocessing
and class order never drift.
Key Metrics
- ~95% classification accuracy on the held-out test set
- 38 disease/healthy classes across 14 crop types (Apple, Corn, Tomato, Potato, Grape, and more)
- ~87,000 training images split 70% train / 15% validation / 15% test
- Single shared
config.pyanddata_pipeline.py— eliminates train/serve preprocessing drift - EarlyStopping and ReduceLROnPlateau callbacks prevent overfitting and wasted compute
- Streamlit inference app with model caching (
st.cache_resource) — model loaded once, not per request - CI via GitHub Actions (Ruff lint + import smoke tests); deployment documented for Streamlit Community Cloud
Portfolio one-liner
Plant Disease Recognition System — End-to-end ML system that classifies plant leaf images into 38 disease/healthy classes using a CNN. Built with TensorFlow/Keras, a single-source config and data pipeline, and a Streamlit web app with correct preprocessing and model caching. Experiment logging and seed control for reproducibility; CI (GitHub Actions) and deployment documented for Streamlit Community Cloud. ~95% accuracy on test; 38 classes across 14 crops.